CoLabs has several spatial transcriptomics and proteomics platforms available for UCSF and external investigators.
Combining our expertise in sample processing, staining, imaging, genomics, and data analysis, CoLabs is positioned to provide multi-disciplinary support for spatial profiling projects. Our labs are located near each other to streamline communication and workflows.
Before you start your project, we recommend that you contact us. We can help you prepare grant materials, design custom panels, and make sure your samples meet requirements. We will also discuss timelines, budgets, and our workflow since our spatial omics instruments are not available for independent use.
Courtesy of the D2B CoLab
We provide guidance on sample preparation as each spatial method has its own sample requirements and slide constraints.
For tissue sectioning and tissue microarrays, we can recommend vendors that we have worked with in the past. In some cases, we may be able to assist with sectioning and TMAs in-house depending on the sample. Please reach out for more information.
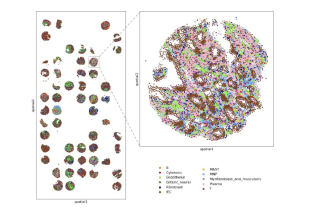
Genomics CoLab offers whole transcriptome analysis through 10x Genomics CytAssist Visium platform. We also have the ability for subcellular resolution on 10x Genomics Xenium platform, with standard and custom designed panels. Compatible samples include both fresh frozen and fixed tissues. Immunofluorescence and H&E staining can be co-visualized on the same sample section.
For custom probes, Genomics CoLab can help you design gene panels from your single-cell RNA sequencing data. Inputs to our Panel DesignR tool include your prioritized gene list, annotated scRNA-seq data, and the option to select certain parameters. Outputs are annotated and ranked panel of candidates, simulated Xenium data, and a file compatible with 10x Genomics’ tool. Please contact us for more information on using this tool.
| Platform: | CytAssist Visium | Xenium |
|---|---|---|
| Detection: | whole genome |
up to 5,000 genes |
| Resolution: | single cell | sub cellular |
| Sample: | fresh frozen, fixed frozen | fresh frozen, formalin-fixed paraffin-embedded |
| H&E and IF staining: | pre-run | post-run |
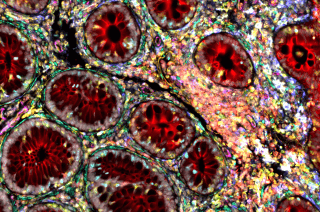
Courtesy of the Kattah Lab
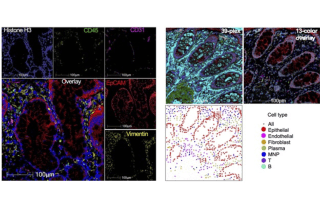
CoLabs offers spatial proteomics analysis using Leica's Cell DIVE Multiplex Imaging system.
| Leica Cell DIVE | |
|---|---|
| Antibody conjugation: | fluorescent dye |
| Panel size: | new markers are constantly being validated, and custom antibody optimization is available |
| Sample: | FFPE tissue only, 4 um thick |
| Compatibility: | H&E |

Courtesy of PALM CoLab
CoLabs can collaborate with you to analyze your spatial data. We have worked on multiple projects and developed pipelines to process, analyze, integrate, and visualize Xenium and CellDIVE datasets. Prior to CellDIVE, we used MIBI and have additional workflows for MIBI data.
Please reach out to discuss how CoLabs can support your computational needs.
1. Who do I contact to start my experiment? Where are you located?
Contact: askCoLabs@ucsf.edu
Location: 513 Parnassus Ave (map)
Medical Sciences Building, 8th floor
2. What if I don’t see the approach that I need for my research? Can CoLabs manage instruments purchased by UCSF labs or programs?
CoLabs is enthusiastic about partnering with UCSF investigators and programs to obtain and manage additional equipment that augment our capacity and bring novel capabilities to UCSF research community. Please contact us to discuss options.